Back to 2025 Abstracts
Scanner-Free Histological Image Acquisition for Artificial Intelligence-Enabled Digital Pathology
Teagan D. Iglesias
*2, Sergii Domanskyi
2, Todd B. Sheridan
1, Javad Noorbakhsh
2, Jeffrey H. Chuang
2, Jill C. Rubinstein
11Hartford Healthcare, Hartford, CT; 2The Jackson Laboratory for Genomic Medicine, Farmington, CT
Background: Histological analysis is the cornerstone of cancer diagnosis and the gold standard for prognostication and treatment planning. In recent years, digitization of histology slides using high-resolution scanners has led to the application of artificial intelligence (AI) algorithms for automated analysis. We and others have demonstrated excellent performance in various tumor classification tasks using such scanned images. However, the high start-up cost for scanner technology creates a disparity in access to this technology. We hypothesize that scanner-free image acquisition using digital cameras mounted on microscopes already present in clinical pathology laboratories provides adequate resolution for our AI pipeline to extract clinically meaningful information.
Study Design: This is a pilot study using BRAF-mutant melanomas at varying stages of targeted treatment with dabrafenib and trametinib, comparing images digitized using a commercial scanner (40x) to those from an 8 megapixel microscope-mounted digital camera (10x, 20x, and 40x objectives). We designed a pipeline for format conversion, automated focus checking, stain normalization, and tissue/background distinction. Preprocessed images are fed forward into our deep learning pipeline for both nuclear segmentation and numerical feature extraction. Logistic regression (LR) is used to fit models for the proof-of-concept task of distinguishing treatment stage: untreated v. early- v. late-treatment.
Results: Images were obtained of glass histology slides from two melanoma models, sampled at various time points during treatment. After quality control, features were extracted from 25 scanned and 76 scanner-free images. Manual inspection by a board-certified pathologist of randomly selected regions of interest confirmed that scanned and scanner-free images are indistinguishable, nuclear segmentation on both image sets could comprehensively identify and quantify cell nuclei, and spatial clustering across scanner-free images could segregate tumor from stromal regions. LR models trained on scanner-free and tested on scanned images successfully distinguished pre-, early-, and late-treatment tumors with precision, accuracy, and recall of 0.93, 0.92, and 0.92, respectively.
Conclusions: We demonstrate that scanner-free image acquisition for AI-enabled digital pathology applications is feasible, providing images of sufficient quality and resolution to allow training of models for tumor classification tasks. Scanner-free images were obtained at a fraction of the cost of using current high-resolution scanners. This pilot provides evidence for the feasibility of implementing digitization workflows with minimal financial and infrastructure investment, establishing that a scanner-free pipeline can mitigate against technology-driven disparities in access to personalized cancer care.
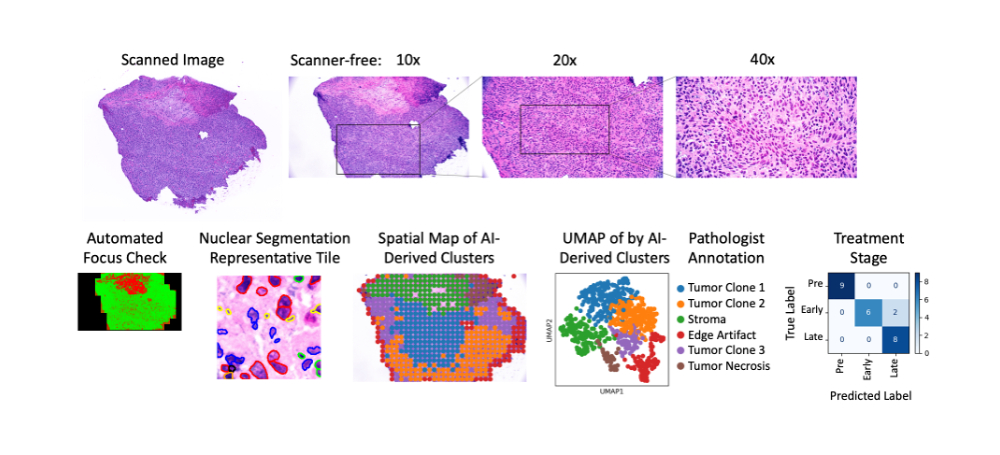
Figure 1. Comparison of scanned versus scanner-free images (top). Bottom row from left to right shows spatial map of automated focus checker (green=in focus), representative image tile with automatically detected cell nuclei, spatial map of clustering by artificial intelligence-derived imaging features, UMAP of AI-derived clusters with pathologist’s manual annotation, confusion matrix showing classification of treatment stage by imaging features.
